Loading Liposomes With Gentamicin to Improve Gentamicin Delivery
Info: 9648 words (39 pages) Dissertation
Published: 9th Dec 2019
Tagged: Pharmacology
AMP Antimicrobial Peptides
BCA Bicinchoninic Acid Protein
CEI Competitive enzyme immunoassay
DLS Dynamic Light Scattering
DSPC 1,2-Distearoyl-sn-glycero-3-phosphocholine
DTX Fluorescence-immunoassay
ELISA Enzyme-linked immunosorbent assay
EMIT Enzyme immunoassay
GLASS Global Antimicrobial Surveillance System
GRAS Generally Regarded as Safe
GUV Giant Unilamellar Vesicles
HPLC High Performance Liquid Chromatography
LFH Lipid Film Hydration
LIA Liposome immunoassay
LUV Large Unilamellar Vesicles
MLV Multilamellar vesicles
MVV Multivesicular vesicles
OLV Oligolamellar vesicles
OPA o-Phthaldialdehyde
PFIA Polarisation fluorescence-immunoassay
SDS-PAGE Sodium dodecyl sulfate polyacrylamide gel electrophoresis
SEM Standard Error of Mean
SUV Small Unilamellar Vesicles
TEM Transmission Electron Microscopy
WHO World Health Organization
- INTRODUCTION
In January 2018, the Global Antimicrobial Surveillance System (GLASS) by WHO showed an increase in antibiotic resistance cases among half a million people with suspected bacterial infections worldwide.1 The report revealed that most of the common yet dangerous infections are resistant to antibiotics; hence, posing as a crucial health and development issue worldwide.1 Unfortunately, development of new antibiotics have decreased over the past 25 years in the pharmaceutical industry due to insufficient federal funding and lack of investment return.2,3 Due to that, improvisation of existing antibiotics are more commonly formulated and optimized.4
In addition, antibiotics that do not adhere to the Lipinski Rule of Five may lead to poor permeability through biological membranes, which tend to be another limiting factor for their effective use. Examples of such antibiotics are the broad spectrum aminoglycoside class of antibiotics that include gentamicin, tobramycin, streptomycin, amikacin and netilimicin. Since they are highly water soluble, these drugs exhibit poor permeability profiles through biological membranes due to the lipid structure of the biological membranes. Also, they exhibit narrow therapeutic window associated with high risks of ototoxicity and nephrotoxicity.5 Due to these factors, liposomal encapsulation of aminoglycoside antibiotics have been researched to enable entry of drug to the intracellular components to enhance antibacterial activity and reduce toxicity.5 Apart from that, a disrupting effect on the bacterial cell membrane could increase the entry of drug into the cell. For this, many ligands have been considered, one of which is the use of antimicrobial peptides (AMPs).6
- LIPOSOMES AS A DRUG DELIVERY SYSTEM
Liposomes have been greatly studied as a drug delivery system with high prospective in the pharmaceutical industry. Due to its amphiphilic properties, they are able to transport small molecule drugs to the site of action especially if poor lipid permeability is a limitation of the drug. Since liposomes are lipid-based carriers, they can permeate through biological membranes to reach target site. Currently, there are many commercialized liposomal formulations that are available in the market such as daunorubicin (DaunoXome®; Gilead Sciences Inc., USA), doxorubicin (Doxil®, Caelyx®; Alza Pharmaceuticals, USA) and amphothericin B (Amphotec®, Ben Venues Laboratory Inc, USA).7,8
2.1 STRUCTURE
In general, a liposome is a lipid bilayer structure that encloses an internal aqueous volume.9 Lipids used for the formation of liposomes is made up of both hydrophilic polar head-group and a hydrophobic non-polar tail-group.9 When added to excess aqueous solutions, these lipids automatically self-assemble such that the hydrophilic head-groups are revealed to the aqueous phase while the hydrophobic tail-groups pair up with each other within the bilayer.9 Thus, liposomes are able to encapsulate hydrophilic and hydrophobic molecules depending on the location of entrapment; hydrophilic drugs may be incorporated in the internal or inter-bilayer aqueous spaces whereas hydrophobic drugs may be incorporated within the membrane itself.9,10
Cholesterol and/or its derivatives are often integrated into the phospholipid membrane to manipulate the mechanical rigidity of phospholipid molecules; leading to compaction, reduced size and improved permeability/stability.10 Cholesterol would arrange themselves within liposomes with hydroxyl groups pointing towards the aqueous phase and the aliphatic chains of cholesterol would line up parallel to the acyl chains of the phospholipid in the bilayer.11 This arrangement would create opposing effects in terms of fluidity, permeability and rigidity based on the temperature it is at (either above or below the phase transition temperature of phospholipids).11 Rigidity is important to ensure minimal leakage from liposomes with maximal stability of liposomal formulations.10
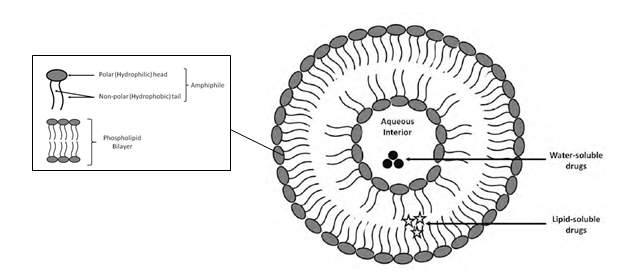
Figure 1: Schematic representation of the structure of liposomes. The compositions of phospholipid bilayers as well as the position of entrapment of drugs formulated into liposomes are displayed (Adapted and modified from Blomme, 2008).12
2.2 CLASSIFICATION
Liposomes are classified based on structure, size, lipid composition, production methods and application as a drug delivery system. All these aforementioned parameters influence the characteristics and in-vitro/in-vivo behaviour of liposomes.15 Due to the height of development and vast research, a detailed and comprehensive review on the classifications of liposomes could not be reported as it is beyond the scope of this report. The following presents a concise outline of the most common classification methods that are relevant to the study conducted.
The most common classification method is based on the structure which eventually influences the size. The size and number of bilayers (lamellae) present affects the circulation half-life and degree of drug encapsulation.13 This classification divides into two groups, i.e.; unilamellar and multilamellar vesicles. Unilamellar vesicles are liposomes which contain a single phospholipid bilayer encapsulating the aqueous core. They can be subdivided into three main groups; small unilamellar vesicles (SUV) which ranges between 0.02 µm to 0.1 µm, large unilamellar vesicles (LUV) which ranges between 0.1 µm to 1 µm (close to the size of living cells) and giant unilamellar vesicles (GUV) which are sized more than 1 µm. On the other hand, multilamellar vesicles (MLV) consist of more than one lamella with sizes from 100 to 1000 nm. Each vesicle is usually made up of three or more concentric lamellae. Oligovesicular vesicles (OVV) are vesicles containing few concentric vesicles inside a much larger one. They are regarded as bilayers and range in sizes from 0.1 µm to 1 µm. Multivesicular vesicles (MVV) consists of two or more vesicles enclosed non-concentrically within a larger vesicle with sizes 0.1 µm and above.13-16
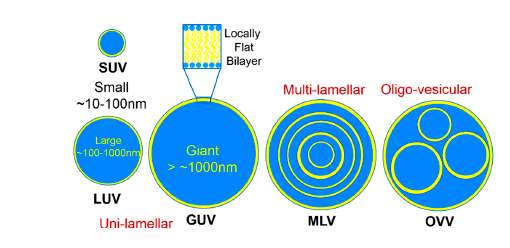
Figure 2: Classification of liposomes according to structural characteristics. (Adapted from Kulkarni C.V., 2016)16
Another common classification is based on production methods of liposome. Methods such as passive/active loading, mechanical/solvent dispersion method and detergent removal method can all be used depending on many criteria such as physicochemical properties of drug that will be entrapped, liposome components (type of lipid and cholesterol used) and the type of medium used to disperse the lipid vesicles (polar or non-polar).14,15,17 Different methods are most likely to produce different structure and sized liposomes as illustrated in Figure 3 below.
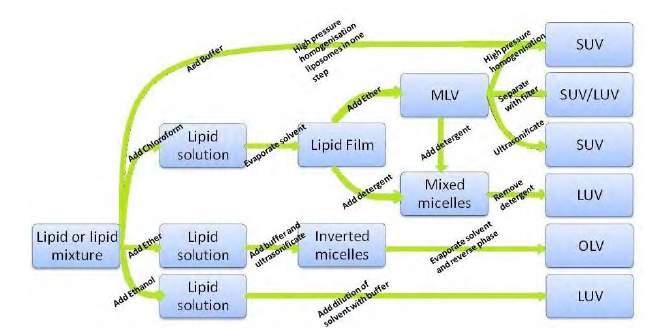
Figure 3: Schematic diagram showing different production methods and the products formed. (Adapted from Müller et al., 1998).
2.3 ADVANTAGES
Since liposomes are made out of non-toxic phospholipids, they are biocompatible, biodegradable and weakly immunogenic. Also, liposomes are able to increase efficacy of drugs that uses it as a carrier due to its ability to permeate across biological membranes as mentioned earlier. Liposomes are able to increase the stability of drugs via encapsulation, which may increase circulation time and half-life of drug. Apart from that, due to the encapsulation, drugs are not able to emit its toxic effects; hence, reducing toxicity of the encapsulated agent. Active targeting and synergistic action may be achieved via coupling with site-specific ligands.14,18 All the above advantages provide the possibility for liposomes to be used to overcome the global threat of antibiotic resistance as mentioned earlier.
2.4 INTERACTION WITH CELLS
Liposome-cell interactions are important as it enables scientists to predict the behaviour of liposomes in-vitro and in-vivo. Many researches on the topic of liposome-cell interactions have been carried out to conclude that liposomes react with cells via various mechanisms such as fusion process, phagocytosis/endocytosis, adsorption and contact release.
Fusion happens when liposomes composed of certain phospholipids link two adjacent cells by fusing with their plasma membrane, thereby linking both the cells. During this linking, liposomal contents are released into the cytoplasm of the cell. 19 However, this does not occur often as liposomes are more often engulfed by endosomes before the fusion process can happen. In this case, liposomes would be transported to lysosomes, and liposome lipids would then be digested by lysosomal enzymes to release the liposomal contents into the cell cytoplasm.20 These processes have been shown to be the result of cell adsorption whereby the liposome attaches to the surface of a cell causing either fusion or endocytosis to occur. This interaction is important as it is the first step before any other liposome-cell interaction occurs.20,21 Another possible interaction is contact release; which is the exchange of liposome and cell bilayer constituents as they are both made up of phospholipids. During this exchange, liposomal contents are released.21
- GENTAMICIN
Gentamicin belongs to the aminoglycoside class of antibiotic widely used to treat severe sepsis. It is sensitive against many Gram negative bacteria such as Pseudomonas aeruginosa, Acinetobacter, Enterobacter and a few Gram positive bacteria such as Staphylococcus.5 This antibiotic causes a bactericidal effect by inhibiting protein synthesis. It does this by tightly binding to 30S and 50S ribosomal subunits to disrupt the integrity of bacterial cell membrane.5 Gentamicin is freely soluble in water and reaches peak serum concentrations in 30-60 minutes upon administration.5 However, it is unable to cross the phospholipid bilayer of biological cell membranes, causing it to be a poorly permeable drug exhibiting poor anti-infective properties. During systemic elimination via kidneys, gentamicin accumulates at the proximal renal tubular cells, leading to mild and reversible nephrotoxicity affecting up to 25% of patients on gentamicin treatment.25 Worse, gentamicin is related to irreversible ototoxicity which is shown to be related to collective lifetime exposure towards it.25
Encapsulation of gentamicin in liposomes done by Swenson et al. showed improved permeability and increased therapeutic index due to increased concentration of gentamicin at intracellular target site.32 Another study by Fierer et al. proved that free gentamicin displayed more toxic side effects in rats as compared to encapsulation of gentamicin in liposomes.33 Multiple methods of drug loading in liposomes have been studied and shown to have various encapsulation efficiencies which led to variable therapeutic effects.6 Hence, the current work done was aimed to elevate the formulation from a usual gentamicin-loaded liposomes (which has already been widely studied) to having another active ingredient adsorbed onto the liposome to improve the overall therapeutic efficacy of the formulation.
One of the challenges that revolve around research regarding gentamicin is the quantification of gentamicin.23 Gentamicin is a complex molecule comprising of three major and a few minor components which poorly absorbs ultraviolet and visible light.22 As this molecule does not contain fluorophores, direct spectrophotometric/fluorometric methods cannot be applied.23 Therefore, detection of gentamicin in liposomes requires some sort of derivatisation and will be discussed further in this paper.24
- NISIN
Nisin, the bacteria-derived peptide used in this study is one of the earliest and most extensively studied lantibiotic.26 It was first isolated in 1947 from Lactococcus lactis by Mattick and Hirsch and has since been used in the food industry; especially in dairy products, meat and canned foods.26,27 Nisin is classified as a bacteriocin and is potent against Gram positive bacteria such as Listeria monocytogenes, Staphylococcus aureus, Bacillus cereus, Lactobacillus plantarum and Micrococcus flavus.28 Stable resistance against nisin is rare due to its dual mechanism of action. Firstly, it is able to bind to lipid II and inhibit the synthesis of peptidoglycan that is crucial for cell membrane structure. Secondly, it forms pores within the cell membrane via interaction with anionic lipids, leading to leakage of cellular constituents and inhibition of cell wall biosynthesis.27-29
Since they can mediate cellular adhesion and invasion in addition to exhibiting their own antimicrobial properties, adsorption of these peptides onto drug-loaded liposomes would utilize the natural mechanism used by these invasive peptides to improve the therapeutic efficiency of an anti-infective formulation.6
Previously, the potential of AMP-adsorbed nanocarriers have been demonstrated by a few studies. Bühler et al. showed that vaccines fused to AMPs adsorbed on microparticles are able to induce strong mucosal immune response.30 Additionally, a study by Autenrieth et al. demonstrated the ability of AMP-adsorbed microparticles to affect the intracellular delivery of vaccines.31 Moreover, Menina et al. researched the ability of gentamicin-loaded, AMP-adsorbed liposomes to target gentamicin to the intracellular components of cells. It showed reduction of infection load by increasing permeability of gentamicin across the cellular membrane and vacuolar compartments. All these mentioned studies used invasin, an AMP encoded by inv gene of Yersinia pseudotuberculosis.6,30,31 So far, there have not been many studies using nisin as both an active ingredient and adjuvant to an antibiotic to improve antimicrobial efficacy against intracellular pathogens. The current work done was aimed to constitute such a study.
One of the obstacle of using nisin and gentamicin in the same formulation is during the quantification of both entities. Although the assays used in this study have been established in current literature, interference of many factors such as amino groups, lipids and temperature of nisin and gentamicin in the assays have not been fully studied. Findings of these interferences will be discussed further in this paper.
In a nutshell, this study was aimed to utilize liposomes as nanocarriers for gentamicin, a poorly permeable antimicrobial with broad spectrum action. Nisin was then adsorbed onto these liposomes to improve the delivery and efficacy of this formulation. Due to time constraints, only physical and chemical characterizations of this formulation were conducted in addition to in-vitro drug release studies.
AIMS
This study was aimed to to load liposomes with gentamicin, a poorly permeable anti-infective and adsorb nisin, an antimicrobial peptide on the liposomal surface to improve gentamicin delivery. Secondly, this formulation was optimized in terms of loading dose and quantification methods. Thirdly, the aim was to characterize the resulting formulation via physical and chemical characterization to enable short-term stability studies and adsorption/encapsulation quantification. Lastly, in-vitro drug release studies were conducted to compare the drug release of gentamicin from gentamicin loaded liposomes and combined liposomes versus release of nisin from nisin adsorbed liposomes and combined liposomes.
HYPOTHESIS
Firstly, loading of liposomes will increase the size (d.nm) and cause reduction in zeta potential (mV) values. In addition, if the loading dose of either nisin or gentamicin is increased, the adsorption/encapsulation efficiency of nisin/gentamicin on liposomes will increase as well. Thirdly, gentamicin will be more difficult to release from combined liposomes compared to the gentamicin loaded liposomes as nisin on the surface will hinder the release of gentamicin. On the other hand, nisin will be released faster compared to gentamicin from combined liposomes as it is adsorbed on the liposome surface as opposed to encapsulated within the liposome.
- MATERIALS
1,2-Distearoyl-sn-glycero-3-phosphocholine (DSPC) purchased from Avanti Polar Lipids Inc., United States. 5-cholesten-3β-ol (>99% cholesterol), Nisin from Lactococcus lactis 2.5%, gentamicin sulphate (meets USP testing specifications, powder), o-phthaldialdehyde 97% (OPA) purchased from Sigma, United Kingdom. 2-mercaptoethanol (>99%) and potassium hydroxide pellets anhydrous >99.97% obtained from Sigma Aldrich, United Kingdom. Bicinchoninic acid protein assay (BCA) reagent was purchased from Thermo Scientific, United Kingdom. Boric acid (analytical reagent grade, powder), chloroform (CHCl3, 99.8%), ethanol (C2H6O, analytical grade reagent) and methanol (CH3OH, >99.9%) were obtained from Fisher Scientific, United States. Phosphate buffer solution (PBS) of pH 7.4, distilled water and deionized water were prepared by Liverpool John Moores University (LJMU) laboratory technicians. All other chemicals and solvents used were at least of analytical grade.
- LIPOSOME PREPARATION
In this study, the preparations of liposomes were performed using lipid film hydration method as described by Bangham and his colleagues.34 In a 250 mL round bottomed flask and in a molar ratio of 1:1 DSPC and cholesterol were dissolved in 5 mL of chloroform: methanol (2:1). The flask was then attached and locked in place using safety clips to a rotary evaporator (Heidolph Laborota 4000 Efficient, Austria) equipped with rotation at 100-150 rpm and a heating bath that was set to 70oC (above the phase transition temperature of DSPC; 55oC) for 1 hour. This led to a formation of a dry lipid film with no traces of organic solvent. The flask was removed and the vacuum outlet of the rotavapor was covered with parafilm to prevent any water evaporation from occurring during the rehydration step. The temperature was lowered to 60oC for this step. 5 mL volume of pre-warmed PBS (pH 7.4) was added to the produced lipid film and the flask was re-attached to the rotavapor to be rotated in the water bath for another hour, leading to the hydration of lipid film and the formation of liposomes. The resulting liposomes were sonicated in an ultrasonic bath (GAH 209 Ultrawave Precision Ultrasonic Cleaning) for 1 hour and then extruded 10 times with 200 nm pore sized polycarbonate membrane filters (Nuclepore track-Etch Extrusion Membrane, Whatman®) in an extruder (Avestin Liposofast LF-50, Germany) that was attached to a water batch (Huber CC3, Germany) at 60oC. Membrane changes were carried out following five extrusions. The final dispersion of liposomes were stored at 4oC.
Initially, unloaded liposomes were produced according to the steps explained above. Loaded liposomes were prepared using the same steps, whereby gentamicin solution of 10 mg/mL loading dose in PBS (pH 7.4) was used instead of PBS (pH 7.4) during the rehydration step. (Figure 1)
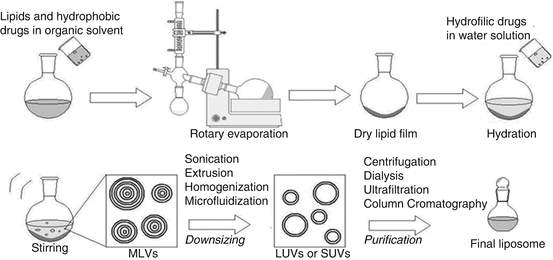
Figure 4: Thin-film hydration method (Adapted from De Arou’ jo Lopez et al.)35
- ADSORPTION OF LIPOSOMES WITH NISIN
A non-covalent corporation of nisin onto the surface of liposomes was performed. Firstly, a nisin stock solution of known concentration was prepared in a 5 mL volumetric flask. Then, 1 mL of nisin stock solution was added to 1 mL of liposomes in a Falcon tube and rotated at 20 rotations/minutes (Multi-Rotator PTR-35 Grant-bio, UK) for 1 hour at room temperature. After an hour, the 2 mL of the resulting formulation was purified to remove residual nisin that were not adsorbed. Next, the ultrafiltrate was collected and the degree of nisin adsorption was then determined by BCA assay.
- LIPOSOME PURIFICATION
As part of the liposomes functionalization procedure and before the analysis of both adsorbed and encapsulated liposomes, liposomal formulations were separated from any residual, non-incorporated components that could affect the chemical characterization. In all cases, the purification process was carried out by centrifugal ultrafiltration (Eppendorf 5810R Centrifuge, Germany) using Centrisart tubes (Centrisart 1, Sartorius, United Kingdom) with a 300, 000 molecular weight cut off membrane (MWCO). This was carried out by first pouring the liposomal suspension into a Centrisart tube, then adding the filtration membrane and centrifuged at 2500 g and 4oC for 30 minutes (acceleration, brake: 9). The ultrafiltrate was then stored separately and the liposomal formulation was made up to volume with fresh PBS. This purification step was repeated twice each time to ensure complete removal of all residual non-liposomal material.
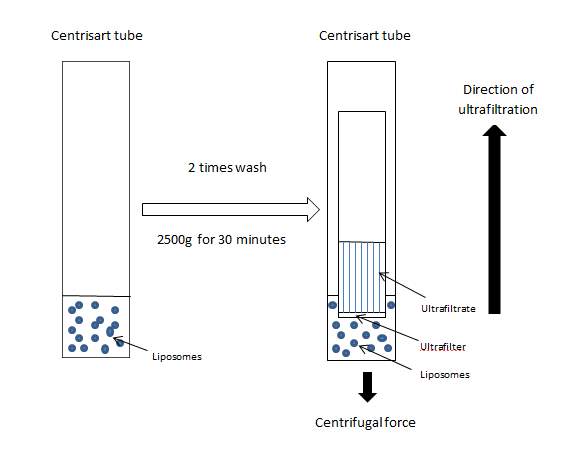
Figure 5: Liposome purification by centrifugal ultrafiltration using Centrisart tubes.
- LIPOSOME CHARACTERIZATION
- Physical Characterization
The size and morphology of liposomes were evaluated to analyze the short term stability of liposomes produced and to determine the behaviour of liposomes in addition to effect of nisin and gentamicin in the liposomal system over time.
- Size and Polydispersity Index (PDI)
The size and PDI were measured using dynamic light scattering (DLS) technique using a Zetasizer (Nano ZS Malvern Instruments, UK). This technique measures Brownian movement to and correlates this to the particle size. Brownian movement is the random movement of particles when it is bombarded by solvent molecules that surround them. The measurement is done by measuring the rate at which the intensity of the scattered light fluctuates when detected using the Zetasizer. 1-in-100 dilution of liposomal formulation was prepared by adding 10 µL to 990 µL of PBS in a clean cuvette. This cuvette was placed correctly into the Zetasizer and the particle size was measured and recorded.
- Zeta Potential
The sample used to measure size and PDI was pipetted out of the cuvette and transferred using a pipette into a capillary cell cuvette. The cell was filled carefully, ensuring no bubbles were formed while filling the cell up to submerge both electrodes in solution. The capillary cell cuvette was then placed into the Zetasizer and the zeta potential (mV) was recorded.
- Transmission Electron Microscopy (TEM)
A copper coated 300-mesh TEM support grid (Agar Scientific Ltd, UK) was immersed in 10 µL of liposomal formulation. The sample was left at room temperature to dry for 10 minutes and the images were then viewed using a transmission electron microscope (MORGAGNI 268 D).
- Chemical Characterization
The development of a liposomal formulation requires careful analysis of key liposomal ingredients to ensure product quality and stability. Two different assays were used to quantify the two active ingredients of this formulation; nisin and gentamicin.
- Nisin Quantification
The amounts of antimicrobial peptide adsorbed to the liposomes were quantified using the bicinchoninic acid protein assay (BCA). This assay incorporates a protein induced biuret reaction with a colorimetric identification of the cuprous cation (Cu+) by bicinchoninic acid. Under alkaline conditions of this assay, a Cu2+ ion is formed and this ion is reduced from Cu2+ to Cu+. It has been revealed that cysteine, cystine, tryptophan, tyrosine and peptide bonds will reduce Cu2+ to Cu1+. Then, two molecules of bicinchoninic acid chelate with one cuprous ion and form a violet coloured product. This bicinchoninic acid-copper complex is water soluble and displays a linear absorbance at 562 nm. The absorbance is directly proportional to the intensity of purple coloured reaction product, and in turn, the protein concentration.
Standard curve dilutions were prepared using a nisin stock solution of known concentration (6 mg/mL or 10 mg/mL) to produce dilutions ranging in concentration from 0 mg/mL to 2.5 mg/mL. The Pierce™ BCA Protein Assay Kit Working Reagent was made by mixing Reagent MA (solution of sodium carbonate, sodium tartrate and sodium bicarbonate in 0.2M NaOH, pH 11.25), Reagent MB (solution of bicinchoninic acid 4% w/v cupric sulphate and pentahydrate solution) and Reagent MC (4% w/v cupric sulphate and pentahydrate solution) in a ratio of 25:24:1. The resultant solution was mixed using a vortex mixer until a resultant homogenous green colour was formed. Then, 150 µL of standard dilutions and dilutions of liposomal formulation was pipetted into each well of the 96-well microplate (Costar Clear Flat Bottom, UK) carefully. Another 150 µL of Pierce™ Working Reagent was added into each well and this mixture was shaken gently on a table top for 30 seconds before being incubated at 37oC (Stuart Incubator S160) for 2 hours. After the incubation period ended, the ultraviolet (UV) absorbance was measured in a plate reader (Softmax Pro 6.5.1) at 562 nm. The amount of nisin adsorbed was then calculated and compared to the standard curve created. This was then expressed as ‘adsorption efficiency’; in which the amount of liposome-bound peptide is given as a percentage from the initial amount of peptide used for adsorption (i.e.; loading dose).36
Equation 1: Adsorption Efficiency6
Adsorption Efficiency (%) = Initial amount of peptide loaded (mg) -Actual amount of peptide (mg) x 100
Initial amount of peptide loaded (mg)
The interference of liposomal lipids in BCA assay was also studied by running a few runs of liposomal formulation that had its lipids broken down by solvents such as 1% ethanol and 1% methanol instead of diluting the liposomal formulations with PBS pH 7.4. The absorbance obtained by these samples was compared with those diluted with only PBS pH 7.4.
- Gentamicin Quantification
A fluorometric procedure developed by Gubernator and his colleagues was used for gentamicin quantification. This method is based on the reaction of primary amine groups of gentamicin with the reagent o-phthaldialdehyde (OPA).23 Under alkaline conditions, this reaction produces a fluorescence that is directly proportional to gentamicin concentration. This method is straightforward, accurate and easy to replicate. This method requires access to a spectrofluorimeter (Softmax Pro 6.5.1) and the samples are read at an excitation wavelength of 344 nm and an emission wavelength of 450 nm.6
To prepare this reagent, 0.4 M boric buffer of pH 10.4 had to first be prepared. Boric acid (0.4 M, 24.7 g) was dissolved in 900 mL of deionized water and the pH was adjusted to 10.4 with 50% (w/v) potassium hydroxide.
OPA reagent was prepared by dissolving 0.2 g of OPA in 1 mL of methanol and 19 mL of 0.4 M boric buffer (pH 10.4) that was prepared beforehand. Then, 0.4 mL of 2-mercaptoethanol (14.3 M) was pipetted in and the pH was adjusted to 10.4 with potassium hydroxide solution. Boric acid and 2-mercaptoethanol were used in order to increase the reaction sensitivity and stabilize the fluorescent product. This reagent was stable for 2-3 days when kept in the dark at 40C.
Standards were prepared using gentamicin stock solution of 0.1 mg/mL ranging in concentration from 0 µg/mL to 40 µg/mL in PBS (pH 7.4). The assay was conducted by adding 0.6 mL of methanol was then mixed with each standard followed by an addition of 0.9 mL of reagent solution (0.1 mL OPA reagent and 0.8 mL methanol).
Prior to gentamicin quantification in liposomal formulation, lipid extraction had to be done due to their interference with this method as they possess primary amino groups.23 To do this, a modified Bligh-Dyer extraction method was used. This was carried out by adding 250 µL of dichloromethane to 200 µL of washed liposomal dispersion. Then, 500 µL of methanol was added and the mixture was vortexed until the solution turned clear. After that, 250 µL of NaOH solution (0.2 M) was added followed by 250 µL of dichloromethane and this was vortexed again. The resulting biphasic system was then centrifuged at 3700 g for 5 minutes in a bench centrifuge. 400 µL of the upper layer was used for gentamicin quantification. This extracted 400 µL was made to volume of 1 mL with PBS (pH 7.4). Then, similar to the method mentioned above, 0.6 mL of methanol was added to each sample, followed by 0.9 mL of reagent solution (0.1 mL OPA reagent and 0.8 mL methanol). This mixture was incubated for 10 minutes in the dark at room temperature before being transferred to an opaque 96-well microplate (Costar Black Flat Bottom, UK) to be read using a microplate reader.
Standards and samples were then incubated for 10 minutes in the dark and the fluorescence was measured in a plate reader (Softmax Pro 6.5.1) with an excitation of 344 nm, emission of 450 nm and low photomultiplier tube (PMT) settings. The amount of gentamicin encapsulated in the liposomes was then quantified by comparing the measured fluorescence of the samples to that of the standard solutions. The encapsulation efficiency is given as a percentage of the initial amount of gentamicin added during liposome preparation (loading dose of 10 mg/mL).6,37
Equation 2: Encapsulation Efficiency 6,37
Encapsulation Efficiency (%) = Actual amount of gentamicin (mg) x 100
Initial amount of gentamicin loaded (mg)
A test to study the interference of nisin in the OPA assay was also conducted. This was carried out by comparing the fluorescence produced by gentamicin stock solution of known concentration compared to a combination of gentamicin and nisin stock solution of equal amount of gentamicin (mg) in it. Then, similar to the method mentioned above, 0.6 mL of methanol was mixed with each sample, followed by 0.9 mL of reagent solution (0.1 mL OPA reagent and 0.8 mL methanol). The sample was incubated in the dark for 10 minutes at room temperature before being transferred to a 96-well microplate (Costar Black Flat Bottom, UK) and read using a microplate reader (Softmax Pro 6.5.1).
- Release Test
In-vitro release of nisin and gentamicin from the liposomes were studied over a period of 24 hours. Centrisart (Sartorius, United Kingdom) tubes with 30, 000 MWCO were filled with 0.5 mL of gentamicin-loaded liposomes, nisin-adsorbed liposomes and a combination of both and made up to 2 mL volume by adding 1.5 mL of PBS and was kept rotating at 35 rpm (Multi-Rotator PTR-35 Grant-bio) at 37oC (Stuart Incubator S160) without its inner tube. At pre-determined time intervals of 15, 70, 135, 200, 265, 330, 395 and 1440 minutes, the inner tube containing the filter is inserted and the entire sample is centrifuged at 2500 g at 4oC for 30 minutes. After each centrifugation, the ultrafiltrate was collected and equal volume of fresh PBS pH 7.4 was added to the release samples to ensure sink conditions are maintained.
The amount of nisin and gentamicin released at each time point were then quantified using BCA and OPA assays, respectively, as previously described (Section 5.2.1 and Section 5.2.2). 3 independent batches consisting of 3 replicates were studied for the combined liposomes whereas 3 independent batches were studied for the gentamicin-loaded liposomes and nisin-adsorbed liposomes to study the intra and inter-variability of data obtained.
Known concentrations of nisin solution (0.5 mg/mL and 1 mg/mL) and gentamicin solutions (30 µg/mL and 40 µg/mL) were incubated and rotated as the liposomal formulations were and release samples at pre-determined time intervals as mentioned above were collected. BCA and OPA assays were carried out to these samples and the absorbance/fluorescence was read using a microplate reader (Softmax Pro 6.5.1). The readings obtained were then compared to that of the freshly prepared stock solution of nisin and gentamicin of the same concentration. This was done to evaluate if temperature, time and extended rotation affects the release study to show a significantly elevated or reduced absorbance/fluorescence reading.
Statistical Analysis
Physical characterization of liposomes
In this study, the liposomes formulated were physically characterized via parameters such as particle size, polydispersity index (PDI), surface charge and morphology. Short term stability testing for these formulations was conducted over a period of 4 days.
- Size (nm), Polydispersity Index (PDI) and Zeta Potential (mV)
1.1.1 Empty Liposomes
In this study, all liposomes were prepared using the lipid film hydration (LFH) method. To prepare empty/unloaded liposomes, the thin film was rehydrated with fresh PBS pH 7.4. Three batches of empty liposomes were prepared before the process of adsorption and encapsulation were done to establish the range of size, PDI and zeta potential obtained. All three batches exhibited sizes ranging slightly lower than 200 nm, which was desirable as cellular uptake is shown to be higher with smaller particle sizes.38
Polydispersity index (PDI) is a parameter used to reflect the size distribution of a sample of nanoparticles. Published studies have shown that the lower the PDI in a formulation, the more homogenous it is, which in turn improves the stability of the formulation. PDI less than 0.3 is considered acceptable to implicate the homogeneity in the formulation studied.39,46 All three of the formulations prepared did not exceed PDI of 0.3, indicating a fairly uniformly distributed formulation.
Zeta potential is a parameter used to measure surface charge and electrostatic stabilisation of a particle. Published literature state that the values between -30 mV and +30 mV indicate an unstable suspension, while values higher than +30mV or lower than -30mV are stable samples as they will be less likely to aggregate.39,40 The batches of empty liposomes prepared showed zeta potential roughly around -5 mV as shown in the figure below. This is indicative of an unstable formulation; hence stability studies were conducted to further analyze this issue. Due to the length of the research, the colloidal stability of liposomes prepared was only monitored over 4 days.
The stability studies conducted showed that over time, the size of liposomes increased from 195.4 nm (+ 3.647 nm) to 205.4 nm (+ 8.163 nm) together with an increment of the PDI from 0.214 (+ 0.063) to 0.335 (+ 0.115). The surface charge became more negative over time, from –5.37 mV (+ 0.521mV) to -7.06 mV (+ 0.582 mV). All values reported in the figure below are the mean of three independent batches prepared.
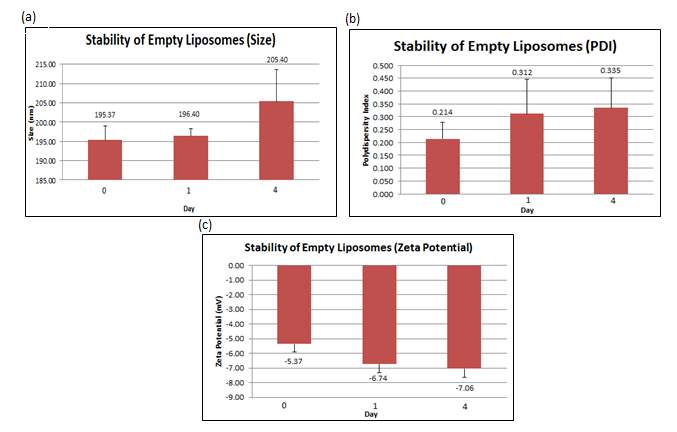
Figure 6: (a) Size, (b) PDI and (c) zeta potential were measured at day 0, 1 and 4 for the empty liposomes. Error bars are representative of the standard error of the mean (SEM) values from three independent preparations.
1.1.2 Gentamicin Loaded Liposomes
Gentamicin loaded liposomes were prepared by hydrating the thin film with a 10 mg/mL gentamicin solution made with PBS pH 7.4. Five independent batches of gentamicin loaded liposomes were prepared and analysed before the combined liposomes were formulated. This is to evaluate and later, to compare the differences between gentamicin loaded liposome with and without surface functionalization. All five batches showed sizes below 200 nm, ranging from 156.4 nm to 183.8 nm. This indicates that loading the liposomes with gentamicin does not notably increase the liposomal size. Not just that, PDI was also shown to be ranging from 0.1 to 0.2 within all five batches, implying homogenous particle sizes in all formulations prepared. Zeta potential on Day 0 was similar to that of empty liposomes and was seen to become more negative after a day, however, as opposed to empty liposomes, the negativity reduced at Day 4.
The stability studies conducted showed that over time, the size of liposomes increased from 168.16 nm (+ 11.168 nm) to 180.28 nm (+ 14.770 nm) together with an increment of the PDI from 0.1706 (+ 0.044) to 0.2246 (+ 0.054). The surface charge became slightly more negative after a day, from –3.11 mV (+ 0.820 mV) to -3.118 mV (+ 0.851 mV). On Day 4, the zeta potential increased from -3.118 mV (+ 0.851 mV) to -2.56 mV (+ 0.719 mv). All values reported in the figure below are the mean of three independent batches prepared.
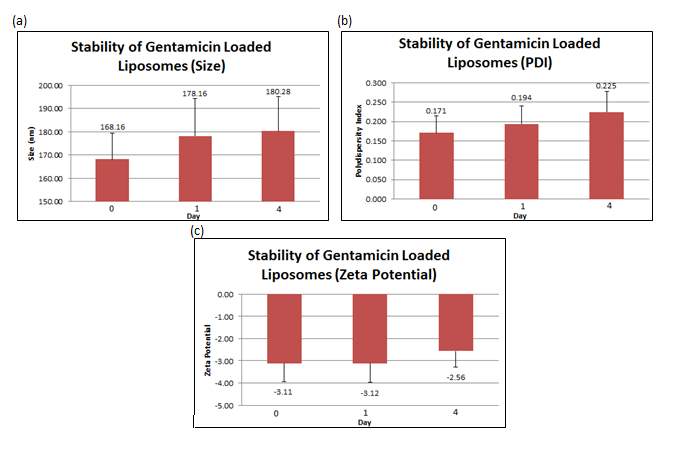
Figure 7: (a) Size, (b) PDI and (c) zeta potential were measured at day 0, 1 and 4 for gentamicin loaded liposomes. Error bars are representative of the standard error of the mean (SEM) values from three independent preparations.
1.1.3 Nisin Adsorbed Liposomes
Nisin adsorbed liposomes were prepared by rotating equal amount of empty liposomes and nisin stock solution of known concentration. In this study, two loading dose concentration were used and compared. 3 independent batches of a 3 mg/mL loading dose were prepared followed by another six independent batches of 5 mg/mL loading dose. All nine of these formulations were physically characterized based on size, PDI and zeta potential.
Liposomes adsorbed with 3 mg/mL nisin solution showed to have slighter smaller sizes ranging from 150.4 nm to 160.8 nm as compared to that of 5 mg/mL which produced liposomes ranging between 183.5 nm to 287.3 nm.
This indicates that the loading concentration of nisin is directly proportional to the size of liposomes produced.
The PDI for the 3 mg/mL formulations remained at a constant 0.2 whereas was more variable with the 5 mg/mL, ranging from 0.2 to 0.3 within all six batches; implying homogenous particle sizes in all formulations prepared. Zeta potential on Day 0 was similar to that of empty liposomes with similar pattern of reduction to become more negative over time for both loading concentrations studied.
The stability studies conducted on the liposomes adsorbed with 3 mg/mL nisin revealed that over time, the size of liposomes increased from 155.56 nm (+ 5.200 nm) to 228.73 nm (+ 72.100 nm) together with an increment of the PDI from 0.227 (+ 0.010) to 0.329 (+ 0.128). The surface charge became more negative over time, from –3.73 mV (+ 0.688 mV) to -4.69 mV (+ 0.604 mV).
In contrast, the stability studies carried out on the liposomes adsorbed with 5 mg/mL nisin showed that over time, the size of liposomes increased from 213.83 nm (+ 39.107 nm) to 238.68 nm (+ 35.000 nm) together with an increment of the PDI from 0.261 (+ 0.071) to 0.329 (+ 0.066). The surface charge became more negative over time, from –3.90 mV (+ 1.236 mV) to -6.80 mV (+ 1.788 mV). All values reported in the figure below are the mean of three independent batches prepared. Values reported in Figure 8 for the 3 mg/mL loading concentrations are the mean of three independent batches whereas the one of 5 mg/mL are the mean of six independent batches.
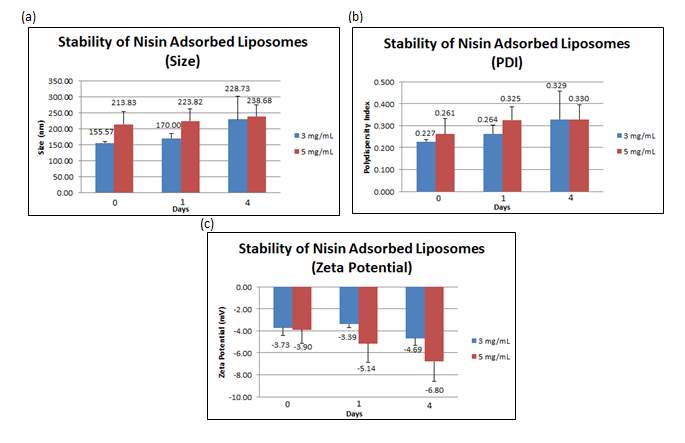
Figure 8: (a) Size, (b) PDI and (c) zeta potential were measured at day 0, 1 and 4 for nisin adsorbed liposomes. Error bars are representative of the standard error of the mean (SEM) values from three/six independent preparations.
1.1.4 Combined Liposomes
The term ‘combined liposomes’ are used in this paper to indicate gentamicin loaded, nisin-adsorbed liposomes. These liposomes were first prepared by loading gentamicin using the thin film hydration method as explained above. Then, the gentamicin loaded liposomes were rotated with nisin stock solution of 5 mg/mL to allow non-covalent bonding. 3 independent batches, each consisting of 3 replicates of combined liposomes were prepared and analysed. All 3 batches showed sizes below 200 nm, ranging from 160.6 nm to 171.5 nm. This indicates that having gentamicin inside the liposomes and nisin on the surface does not significantly affect the size of the liposomes. Not just that, PDI was also shown to be ranging from 0.1 to 0.2 within all 3 batches, implying homogenous particle sizes in all formulations prepared. Zeta potential on Day 0 was similar to that of empty liposomes and was similarly seen to become more negative over time.
The stability studies conducted showed that over time, the size of liposomes increased from 169.3 nm (+ 3.811 nm) to 233.7 nm (+ 38.759 nm) together with an increment of the PDI from 0.166 (+ 0.043) to 0.316 (+ 0.021). The surface charge became slightly more negative after a day, from –3.73 mV (+ 0.196 mV) to -5.70 mV (+ 0.925 mV). All values reported in the figure below are the mean of three independent batches consisting of 3 replicates each.
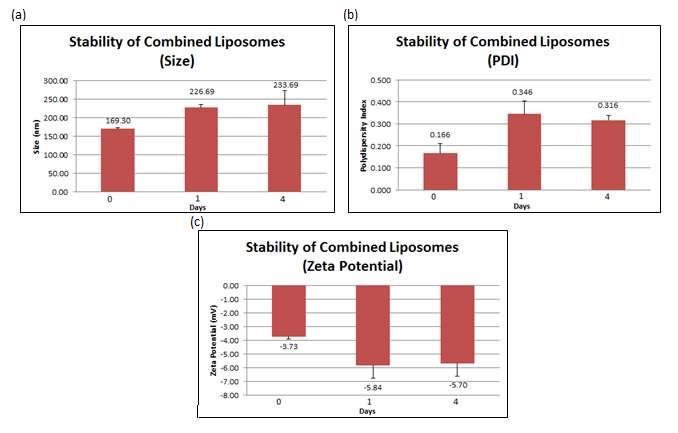
Figure 8: (a) Size, (b) PDI and (c) zeta potential were measured at day 0, 1 and 4 for combined liposomes. Error bars are representative of the standard error of the mean (SEM) values from three independent preparations.
1.2 Liposome Imaging (TEM)
The various liposomes formulated were then imaged using Transmission Electron Microscopy (TEM) to characterize its size and shape. TEM is useful as it can directly visualize single particles and sometimes, the inner structure such as number of lamellae.41 The images obtained show a mixture of spherical and aspherical liposomes at approximately 200 nm in diameter. This size visualized tally with the results obtained via DLS reported earlier. More detailed information regarding the structure was unable to be visualized with or without staining.
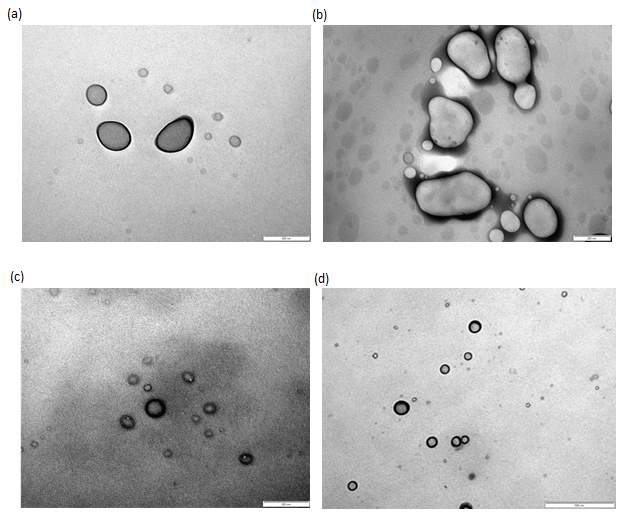
Figure 9: TEM images without staining of (a) empty liposomes (200x magnification), (b) gentamicin loaded liposomes (200x magnification), (c) nisin adsorbed liposomes (500x magnification) and (d) combined liposomes (500x magnification).
- Chemical Characterization of Liposomes
In this study, the liposomes formulated were chemically characterized by evaluating the encapsulation and adsorption efficiency of the gentamicin and nisin, respectively. Assays selected for quantification methods were decided upon reviewing published literature for the most suitable option in terms of time, cost and accuracy.
2.1 Encapsulation and Adsorption Efficiency
- Encapsulation of Gentamicin
Encapsulation efficiency is the amount of gentamicin that has successfully been encapsulated into the liposomes versus the initial loading amount of gentamicin used in the preparation. This quantification was measured via OPA assay which emitted fluorescence that is directly proportional to gentamicin concentration. 6 independent batches of gentamicin loaded liposomes and 3 independent batches of combined liposomes that consist of 3 replicates each were prepared and quantified. This showed an average of 27.91% (+ 2.031%) encapsulation efficiency in gentamicin loaded liposomes compared to 10.93% (+ 2.454 %) of encapsulation efficiency in combined liposomes, both quantified a day after gentamicin loading (Figure 10).
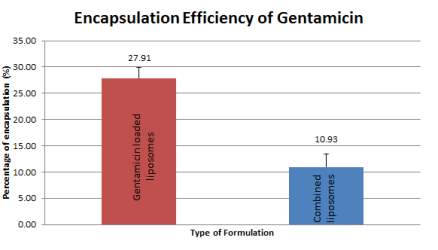
Figure 10: Encapsulation efficiency of gentamicin from gentamicin loaded liposomes and combined liposomes at Day 1 of preparation. Error bars are representative of the SEM values from six/three independent batches.
Three variables were studied to determine the effect of lipid content, the presence of nisin and elevated temperatures on gentamicin quantification calculated based on OPA assay. Results obtained show that lipid content and elevated temperatures affected the accuracy of OPA assay conducted whereas the presence of nisin in the formulation did not affect the results of the gentamicin quantification.
- Adsorption of Nisin
Adsorption of nisin refers to the amount of nisin that has been functionalized on the surface of the liposome in relation to the initial loading amount of nisin. This quantification was measured via BCA assay in which the absorbance value produced is directly proportional to the nisin concentration. In this study, the adsorption efficiency of nisin was quantified in two loading concentrations; 3 mg/mL and 5 mg/mL which showed adsorption efficiency of 15.59% (+2.596%) and 21.94% (+1.576%). As expected, the higher the loading concentration, the higher the adsorption efficiency.
Due to better adsorption efficiency, 5 mg/mL loading concentration was selected to be used in the combined liposomes formulation. The presence of gentamicin in the liposomes seem to not have an effect on the adsorption efficiency of nisin as the mean value of nisin adsorption in combined liposomes were 21.93% (+2.709%).
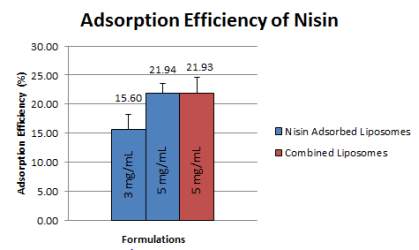
Figure 11: Adsorption efficiency from nisin adsorbed liposomes and combined liposomes at Day 0 of preparation. Error bars are representative of the SEM values from four/five/three independent batches, respectively.
Interference of lipid and elevated temperatures towards BCA assay was studied in this research. It appeared that lipids did not affect this study as the absorbance values remained the same for lipid extracted samples and non- lipid extracted samples. However, when samples were left out for a prolonged time at 37oC, it was found that the absorbance values increased, leading to falsely elevated nisin quantification.
3. In-Vitro Drug Release test
– Gentamicin Loaded liposomes
– Nisin Adsorbed liposomes
– Combined liposomes
– comparison of nisin only vs combined
-comparison of gentamicin only vs combined
– comparison of nisin vs genta
– comparison of all
- Physical Characterization of Liposomes
Liposomes were first loaded with gentamicin back in 1980 by Morgan and Williams using phosphatidylcholine, phosphatidylserine, stearylamine and cholesterol in variable molar ratios to study the different types of liposomes formed.42
Previous studies show liposomes prepared via LFH method using DSPC/cholesterol exhibit mean sizes ranging from 90 nm to 175 nm after being extruded through filters of 100 nm pore size.43,44 This is more or less consistent with the results obtained that show mean sizes ranging from 155.57 nm (+ 5.200 nm) to 238.68 nm (+ 35.000 nm) for all formulations prepared in this study. In addition, there was only a slight percentage of increment in liposome size from Day 0 to Day 4 for empty liposomes (5.134%), gentamicin loaded liposomes (7.207 %) and nisin adsorbed liposomes (11.621 %). At most, there was a 38% liposome size increment from Day 0 to Day 4 shown by combined liposomes.
Additionally, studies by Laridi et al. show that the interaction and binding between cationic nisin and anionic lipid causes rearrangement of particles which in turn, reduces liposome particle size.48 However, this was not observed in this study as empty liposomes were seen to have a mean size of 195.37 nm (+ 3.647 nm) whereas nisin adsorbed liposomes (5 mg/mL) had mean size of 213.83 nm (+ 39.107 nm).
Other studies revolving around liposomal formulations made of DSPC/cholesterol show PDI less than 0.2. The formulations prepared in this study showed PDI ranging from 0.1 to 0.2, indicating fairly monodisperse particle sizes in the formulations prepared. However, all formulations in this study showed slight increment in PDI over time, up to slightly above 0.3.
Particles with with repulsive charges (negative zeta potential values) behave as an energy barrier to control the stability of dispersion and deflecting the attraction between particles to avoid aggregation.44 Published literature reports zeta potential of DSPC/cholesterol liposomes ranging from -1 mV to -3 mV. This low negative charge is similar to the findings of this study which reports zeta potential values ranging from -3 mV to -5 mV. With time, they became more negative, but never achieving the stable range of -30 mV to +30 mV desired by pharmaceutical formulations.39,40 Moreover, it was also seen that zeta potential became slightly less negative upon adsorption of nisin. A study by Vukomanovic et al. explained that the cationic nature of nisin causes this reduction in negative surface charge.49
Due to the zeta potential being in the unstable range, crosslinking between lipids and/or electrostatic interactions between the liposomes and unbound nisin/gentamicin could have caused agglomeration which may be a probable cause of increment in particle size and PDI over time as mentioned above. Studies have shown that incorporating polyethylene glycol (PEG)-lipid could increase the surface negative charges from -1 mV to -13 mV to enhance stability.44,46,47 Moreover, in 1985, Straubinger et al. reported considerable evidence to indicate that anionic liposomes are better compared to neutral and cationic liposomes in terms of functional delivery of therapeutic molecules to intracellular compartments of cells.20
Based on the results obtained, all formulations prepared were able to successfully maintain their colloidal characteristics over 4 days with storage at 4oC. The presence of two saturated fatty chains with 18 carbon atoms in DSPC increases rigidity by reducing the freedom of movement of lipophilic chains. This most likely led to lower drug-lipid interaction which enhanced the stability of the formulations prepared.44,45
- Chemical Characterization of Liposomes
Two literatures have highlighted the relationship between types of lipid used and encapsulation/adsorption efficiency. It was seen that lipids with higher lipid composition has a significant effect on the incorporation efficiency. Both studies show DSPC having better encapsulation/adsorption efficiencies compared to other lipids.50,51
A study by Menina et al. showed 50% reduction in encapsulation efficiency of gentamicin after 3 washes and suggested that the extent of this reduction could be minimized by reducing the washing steps.6 In this study, the encapsulation of gentamicin was quantified with and without nisin on the surface. The results show a reduction from 27.91% (+2.031%) to 10.93% (+2.454%), which is 60.83% reduction in encapsulation efficiency. Although this could be explained by the addition of washing steps during adsorption procedure, it shows that simply reducing the number of washing steps may not be sufficient to minimize the extend of this reduction. As mentioned in 2.1.1, lipid content showed intense method interference in the OPA assay. This is due to the presence of amino groups in lipids.23 Therefore, lipid extraction using a modified Bligh-Dyer method was carried out prior all quantification studies. Apart from that, since elevated temperatures also showed interference in OPA assay, all assay procedures were conducted under room temperature.
Due to time constraints, loading efficiency of liposomes could not be calculated as the actual amounts of lipid components were not quantified. The amounts of phospholipids and cholesterol could be quantified using Stewart assay and High Performance Liquid Chromatography (HPLC), respectively.6
In addition, quantification results based on OPA assay could not be confirmed via other techniques due to limited time. Several methods of measuring gentamicin in liposomes have been established, such as polarisation fluorescence-immunoassay (PFIA), enzyme immunosorbent assay (ELISA), enzyme immunoassay (EMIT), fluorescence-immunoassay (DTX), colorimetric methods, microbiological methods, fluorometric assay and liposome immunoassay (LIA).23
As mentioned in 2.1.2, higher loading concentration has been associated with higher adsorption efficiencies.16 The increase from 15.60% (+2.596%) to 21.94% (+1.576) proves the aforementioned statement. A study by Chibowski and Szcześ demonstrated that when both the liposome and adsorbing molecule possess small negative zeta potentials, hydrogen bonding and London dispersion forces are more likely to be the driving force for adsorption as opposed to electrostatic interactions.52 This could be why the presence of gentamicin in liposome did not affect the adsorption efficiency of nisin in this study.
For calculations, indirect method was used to calculate the adsorption efficiency of nisin, i.e.; the assumption made was that any amount of nisin not left in the ultrafiltrate was encapsulated within the liposome (refer Equation 1 in 5.2.1). This may not be accurate as some nisin could have been stuck on the Centrisart tube walls or on the ultrafilter itself. Thus, confirmatory tests should be done in order to confirm the results of the peptide quantification by BCA assay. A study published by Menina et al. performed sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) to confirm the results of BCA quantification.6 Other methods that may be used include HPLC, ELISA and competitive enzyme immunoassay (CEI).48,53
Cite This Work
To export a reference to this article please select a referencing stye below:
Related Services
View allRelated Content
All TagsContent relating to: "Pharmacology"
Pharmacology involves the study of drugs and how they affect the body. A pharmacologist contributes to drug development by researching and testing how the body reacts to medication, and whether the medication can have a positive impact on the body in terms of fighting illness and disease.
Related Articles
DMCA / Removal Request
If you are the original writer of this dissertation and no longer wish to have your work published on the UKDiss.com website then please:




